Services


The UNLV Genomics Core Facility offers full service Affymetrix GeneChip® microarray processing of Gene 1.0 ST expression arrays. We also offer processing of RNA extractions and have a bioinformatics specialist on site for data analysis. The UNLV Genomics Core Facility is capable of processing 3'IVT, Exon, miRNA, SNP, Tiling and other Affymetrix array types. Because microarrays are an individualized service, please contact the UNLV Genomics Core Facility Manager Casey Hall-Wheeler to discuss your specific needs.
For Gene and Exon expression array services, the user provides total RNA sample and user provides the chip. Listed prices include: front-end QC analysis of RNA sample using Agilent Bioanalyzer, two-cycle cDNA synthesis, labeling, and Affymetrix GeneChip® Instrument System for hybridization, washing, staining and scanning.
Rates for Full-Service Affymetrix GeneChip® Processing
- Gene 1.0 ST Array (user provides chip) = $390/sample
- Exon 1.0 ST Array (user provides chip) = $500/sample
- Other Arrays (Tiling, miRNA, SNP arrays, etc.) = Inquire
Reduced pricing is available for processing large array orders. Please contact the UNLV Genomics Core Manager, Casey Hall-Wheeler for more information.

The UNLV Genomics Core Facility offers full service flow cytometry. Flow cytometry provides quantitative measurements of heterogeneous cell populations based on antigen expression or other molecular probes. Flow cytometry uses light scatter and fluorescence intensity signals to generate specific multi-parameter data of cell populations and particles which we analyze using our BD CellQuest™, ModFit LT™, and FlowJo™ software. The BD FACSCalibur flow cytometer at the UNLV Genomics Core Facility is equipped with two lasers (488 nm and 635 nm) and six detectors. The four fluorescence detectors monitor in the green, orange, red, and dark red regions of the spectrum. The interception of cells with the laser beam produces both forward and side light scatter, which is captured by the FSC/SSC detectors and used to obtain information regarding particle size and intrinsic properties.
Applications
- DNA content (cell cycle, proliferation, ploidy)
- Protein expression, localization, modifications
- Immunophenotyping (cell surface antigens, CD markers)
- Intracellular antigens (various cytokines, secondary mediators, enzymatic activity)
- Intracellular ionized calcium, magnesium, pH
- Apoptosis (quantification, DNA degradation, mitochondrial membrane potential, permeability changes, caspase activity, cell viability)
Flow Cytometry: Examples of our Data
For more information, including consultation for fluorochrome combination feasibility, necessary controls prior to designing your experiment, obtaining protocols, and scheduling flow cytometry, please contact the UNLV Genomics Core Facility Flow Cytometry Technician, Casey Hall-Wheeler.
Rate for full service flow cytometry = $45/hour. Selected flow cytometry reagents available at cost. We currently provide hundreds of single or multicolor conjugated antibodies for pilot studies at no charge.

The Agilent 2100 Bioanalyzer utilizes automated gel electrophoresis and microfluidics technology to separate, stain, destain, detect, image and analyze RNA, DNA and protein samples. This platform can be used to determine the size, quantity and quality of RNA, DNA and protein as well as two color flow cytometry of cells. This instrument offers a wide range of applications, including:
- RNA integrity (RIN) and quality control for microarrays and RT-PCR
- Separate, verify and analyze miRNA after extraction procedures
- Quantification and size determination of PCR fragments and restriction enzyme digests
- Quality assessment, sizing and quantification of fragmented DNA or DNA sequencing libraries for Next-Generation Sequencing
- Protein expression and purification
- QA/QC of antibodies
- QC purity of mtPCR before sequencing
- Microsatellite and PCR-RFLP
- Protein profiling
The Agilent 2100 Bioanalyzer can also be utilized to run simple automated two color flow cytometry applications including:
- Optimize siRNA transfections for gene silencing by evaluating cell viability and delivery efficacy with fluorescent probes
- Detect GFP expression levels
- Detect non-fluorescent intracellular and cell surface protein expression through antibody staining
- Detect apoptosis using Annexin V/Calcein staining
Application Details/Samples of Data
The UNLV Genomics Core Facility currently offers full service RNA analysis using the Agilent RNA Nano 6000 reagents and chips. These chips have the capacity to run up to 12 RNA samples per chip, require 1 µl input volume, have a quantitative range of 25-500 ng/µl and a qualitative range of 5-500 ng/µl.
Rate for full service RNA analysis = $53.41/chip (chip included in price)
We also offer full service for all other Agilent RNA, DNA, protein and flow cytometry chips. Please contact the UNLV Genomics Core Facility Manager Casey Hall-Wheeler for pricing on all other Bioanalyzer applications.
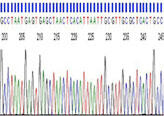
The UNLV Genomics Core Facility offers Sanger DNA sequencing services. Sanger sequencing is run using an either an ABI 3130 or ABI 3500 Dx genetic analyzer and ABI BigDye v3.1 Terminator sequencing chemistry. The typical contiguous read length is > 900 bp, with QV's >20 for high quality, clean DNA templates. The typical turn-around time is two to four business days, although this may change depending on our sample workload.
Sample Submission Information
To submit samples for sequencing, please provide the appropriate amount of DNA template (see the chart below) and primer (3.2 pmol to 10 pmol) in 0.2 ml flat-cap PCR tubes (no strip tubes please) in a final volume of 10 ul of nuclease-free water. Please do not submit templates in TE buffer for the EDTA will chelate the metal ions (Mg++) required for the cycle sequencing reaction.
| Template Type | Amount Required |
|---|---|
| PCR product: 100 to 200 bp | 1 to 3 ng |
| PCR product: 200 to 500 bp | 3 to 10 ng |
| PCR product: 500 to 1000 bp | 5 to 20 ng |
| PCR product: 1000 bp to 2000 bp | 10 to 40 ng |
| PCR product: >2000 bp | 40 to 100 ng |
| Bisulfite converted genomic DNA-PCR product | 3 to 10 ng |
| Single-stranded DNA | 50 to 100 ng |
| Double-stranded DNA | 200 to 500 ng |
| Cosmids, BAC | 0.5 to 1.0 ug |
| Bacterial genomic DNA | 2 to 3 ug |
Submitted samples must be clearly labeled in 0.2 ml flat-cap PCR tubes and each sample must have a unique identifier. The requestor must complete a DNA Sequencing Sample Submission Form and send it by email to Casey Hall-Wheeler at: casey.hall@unlv.edu before bringing sequencing samples to the Genomics Core.
Measures must be taken by the investigator to provide the cleanest DNA template possible for the overall success of the sequencing reaction and data is highly dependent on the quality and quantity of the DNA sample. A pGEM 3Zf(+) control is run with every sample set to ensure instrument and technical performance. Unincorporated dye terminators are removed from the extension products with using the BigDye XTerminator Purification Kit. We can use specialized cycle sequencing and genetic analyzer run protocols for difficult to sequence templates including short PCR products, GC-rich templates, or sequencing primers with non-standard annealing temperatures (inquire for details).
We run sequencing Monday through Thursday by scheduled appointment only. Our turn around time is between two to four days from the time you submit your samples until you have data in hand, although this timing is dependent on our current workload. To schedule an appointment for DNA sequencing services, please call Casey Hall-Wheeler at the UNLV Genomics Core at 702-895-1057.
Universal Sequencing Primers Offered by the UNLV Genomics Core
The UNLV Genomics Core offers the following sequencing primers: T7 promoter, T7 terminator, SP6, pBABE3', pBABE5', pGEX3', pGEX5', CMV forward and M13 forward and reverse. The sequences for these primers are:
| Primer Name | Sequence |
|---|---|
| T7 Promoter | 5'-TAATACGACTCACTATAGGG-3' |
| T7 Terminator | 5'-GCTAGTTATTGCTCAGCGG-3' |
| SP6 | 5'- GATTTAGGTGACACTATAG -3' |
| pGEX-3-prime | 5'-CCGGGAGCTGCATGTGTCAGAGG-3' |
| pGEX-5-prime | 5'-GGGCTGGCAAGCCACGTTTGGTG-3' |
| pBABE-3-prime | 5'-ACCCTAACTGACACACATTCC-3' |
| pBABE-5-prime | 5'-CTTTATCCAGCCCTCAC-3' |
| CMV-fwd | 5'-CGCAAATGGGCGGTAGGCGTG-3' |
| M13 reverse(-24) | 5' -AACAGCTATGACCATG-3' |
| (-21) M13 Forward | 5´-TGTAAAACGACGGCCAGT-3´ |
Cost for DNA Sequencing
The fee for DNA sequencing for samples that are ready to run and do not need to be manipulated by us is $6.25 per sequencing reaction. The fee for DNA sequencing for samples that need to be manipulated by us (addition of primer, verify template concentration on the Nanodrop, transfer submitted samples that are not in 0.2 ml flat-cap PCR tubes into 0.2 ml flat-cap PCR tubes) is $7.50 per sequencing reaction. For a complete list of our DNA sequencing pricing, please see our Fees page.
More information regarding our DNA sequencing services can be found at the following links:
The UNLV Genomics Core Facility offers full service sample processing using the Qubit® 3.0 Fluorometer. The Qubit® 3.0 Fluorometer accurately measures dsDNA, oligos, RNA, microRNA, and protein at low concentrations, using as little as 1 uL of sample. The Qubit® 3.0 is more sensitive than UV absorbance, as it uses highly sensitive Qubit® quantitation assays containing fluorescent dyes that emit a signal only when bound to the target molecule. This specificity minimizes the effects of contaminants such as free nucleotides, intact/degraded DNA and RNA, protein, and excess salts. The Qubit® 3.0 is ideal to quantify samples used in downstream applications such as quantitative real-time PCR or next-gen sequencing. The Qubit® 3.0 can also be used as a mini-fluorometer by selecting either the blue LED excitation source with emission in the green and far-red channels, or the red LED excitation source with emission the far-red channel, to generate RFU values in the 300 - 1,000 nm range.
The Qubit® Assay Kits available for sample processing services in the Genomics Core are:
- Qubit® dsDNA BR Assay Kit – used for accurate quantitation of double-stranded DNA (dsDNA) with initial sample concentrations between 100 pg/µL and 1000 ng/µL. This kit is highly selective for dsDNA over RNA and can quantify sample volumes between 1 µl and 20 µl. Common contaminants such as salts, free nucleotides, solvents, detergents, or protein are well tolerated in the assay.
- Qubit® dsDNA HS Assay Kit – used for accurate quantitation of double-stranded DNA (dsDNA) with initial sample concentrations between 10 pg/µL and 100 ng/µL. The assay is highly selective for dsDNA over RNA and can quantify sample volumes between 1 µl and 20 µl. Common contaminants, such as salts, free nucleotides, solvents, detergents, or protein are well tolerated in the assay.
- Qubit® RNA HS Assay Kit – used for accurate quantitation of RNA with initial sample concentrations between 250 pg/µL and 100 ng/µL in sample volumes between 1 µl and 20 µl. The assay is highly selective for RNA and will not quantitate DNA, protein, or free nucleotides. Common contaminants, such as salts, free nucleotides, solvents, detergents, or protein, are well-tolerated in the assay.
- Qubit® Protein Assay Kit – used for accurate quantitation of protein samples ranging from 12.5 µg⁄ml to 5 mg⁄ml in sample volumes between 1 µl and 20 µl. This assay is highly selective for proteins, exhibits low protein to protein variation, and is accurate in the presence of reducing reagents, but not in the presence of a large amount of detergent. Common contaminants, such as reducing reagents (DTT, β-mercaptoethanol), salts, free nucleotides, amino acids, solvents, or DNA, but not detergents, are well tolerated in the assay.
The rate for full service Qubit® 3.0 Fluorometer sample processing = $3.00/sample, which includes the Qubit® reagents, consumables, sample processing, and electronic data delivery. Alternatively, labs who wish to quantify their own samples can purchase their own Qubit® Assay Kit and can use the Qubit® 3.0 Fluorometer free of charge.
To schedule an appointment for Qubit® sample processing services, or for training on the Qubit® 3.0 Fluorometer, please contact the UNLV Genomics Core Facility Manager, Casey Hall-Wheeler.
Equipment

The Bio-Rad CFX96 Touch™ Real-Time PCR Detection System features an integrated touch screen which allows users to set up runs quickly and monitor amplification traces in real time. This system can multiplex up to five different targets using as low as 10 uL reaction volume. The CFX96 Touch™ uses six filtered LEDs for fluorophore excitation and six filtered photodiodes to detect fluorescence emission, with an excitation range of 450-730 nm. The CFX96 Touch™ has a programmable thermal gradient, melt-curve function, and can detect as low as one copy of target sequence in human genomic DNA with a dynamic range of 10 orders of magnitude. The CFX Manager™ Software offers advanced data analysis tools for normalized gene expression, allelic discrimination, end-point analysis, and analysis of Bio-Rad PrimePCR™ Assays and Panels.

The Bio-Rad iCycler iQ™ Real-Time PCR Detection System can simultaneously measure up to four different fluorophores in each tube of a 96-well PCR plate. This system offers an excitation range of 400-700 nm, customizable filter wheels, programmable temperature gradient, as well as melt-curve and allelic discrimination analysis. The iQ™ optical system software v3.1 is available for RT-PCR product analysis.
The Qubit® 3.0 Fluorometer accurately measures dsDNA, oligos, RNA, microRNA, and protein at low concentrations, using as little as 1 uL of sample. The Qubit® 3.0 is more sensitive than UV absorbance, as it uses highly sensitive Qubit® quantitation assays containing fluorescent dyes that emit a signal only when bound to the target molecule. This specificity minimizes the effects of contaminants such as free nucleotides, intact/degraded DNA and RNA, protein, and excess salts. The Qubit® 3.0 is ideal to quantify samples used in downstream applications such as quantitative real-time PCR or next-gen sequencing. The Qubit® 3.0 can also be used as a mini-fluorometer to generate RFU values in the 300 - 1,000 nm range by selecting either the blue LED excitation source with emission in the green and far-red channels, or the red LED excitation source with emission the far-red channel.
The Amersham Typhoon 5 Biomolecular Imager is a versatile scanner that can image multi-fluorescent, radioisotope-labeled, phosphorescent, chemiluminescent, and colorimetric samples on gels, membranes, phosphor screens, multi-well plates, culture dishes, glass slides, and tissue sections. Fluorescence scans using 5 laser excitation sources, blue 488nm; green 532nm; red 635nm; IR short 685nm; IR long 785nm, facilitate multiplexing and visualization of all visible and IR fluorescently-conjugated antibodies and labels. Accurate quantitation with signal detection as low as 3 pg of protein and differences across a dynamic range with greater than five orders of magnitude. Pixel resolution as low as 10 µm produces high-resolution images. The large scanning area of 40 × 46 cm supports high sample throughput, enabling simultaneous imaging of up to 20 gels or blots measuring 10 × 8 cm in size, or up to 9 multi-well plates in a single scan. The UNLV Genomics Core offers IQTL software for image analysis of relative band intensity and purity analysis of gels/blots, colony counting, and relative fluorescence in plates.
The Shimadzu MALDI-TOF 8020 is a benchtop linear mode (positive-ion) mass spectrometer that provides rapid mass measurement of proteins, peptides, and other biological and organic samples, utilizing a 200 Hz solid-state laser (355 nm) and MALDI Solutions Data Acquisition software for sample acquisition and data analysis.

The DigiLab Genomic Solutions® OmniGrid 100 Microarrayer is a high capacity multi-axis microarrayer that prints DNA samples onto solid substrates. This instrument is fully automated and is capable of handling of up to 72 input plates and holds 100 standard 1" x 3" glass slides. This instrument possesses a 48-pin head that can print in excess of 100,000 features per slide with a resolution of 2.5 µm on the X and Y-axis and 1.25 µm on the Z-axis. The samples are spotted onto the slides in patterns of rows and columns known as sample groups that can be adjusted according to personal preference.

The GenePix® 4000B Scanner can be used to scan DNA and protein microarrays as well as tissue and cell arrays. This scanner has dual lasers that can acquire data at wavelengths of 532 nm (Cy 3) and 635 nm (Cy 5) simultaneously. This instrument scans standard 1" x 3" glass slides and produces images with a resolution range of 5 µm to 100 µm. GenePix® Pro software is available for microarray image acquisition and data analysis, which uses industry standard GAL and GPR file formats.

The Nanodrop 1000 is a full spectrum spectrophotometer (UV and visible spectrum, 220-750 nm) that can measure the absorbance of DNA, RNA, protein and dyes using as little as 1 µl sample volume without the use of cuvettes or capillaries. This instrument is routinely used to quantify and check the purity of RNA and DNA before proceeding with PCR or microarray assays.
The SpectraMax Plus384 is a full spectrum spectrophotometer (190-1000 nm) which contains both a cuvette port and a microplate drawer. Any standard cuvette, 12 x 75 mm test tube, 96 or 384-well microplate can be used to obtain absorbance measurements.

Hyb4 Microarray Hybridization System
The Hyb4 is a microarray hybridization system that functions to automate thermal and wash cycles for hybridization of microarrays spotted on standard glass slides.
The Beckman Coulter Avanti J-E centrifuge is a refrigerated floor model preparative centrifuge. The micro-processor controlled Avanti J-E can be programmed to run at temperatures between 40°C and -10°C. Available rotors include the JA-25.50 rotor with a maximum speed of 21,000 rpm, and the JA-10 rotor with a maximum speed of 10,000 rpm.
The UltraPro 5000 Cryostat is a temperature controlled system used to prepare frozen sections of biological tissues for microscopic examination. This instrument includes a 5040 rotary retracting microtome and allows for tissue orientation in the x and y plane, as well as antiroll plate angle adjustment.
The Eppendorf Mastercycler gradient is a thermal cycler designed for the amplification of nucleic acids. This thermal cycler possesses a programmable heated lid and gradient function, which allows the entire block temperature to be varied within a 20°C range. Peltier elements control rapid heating speeds up to 3°C per second and cooling speeds up to 2°C per second in a temperature range of 4°C to 99°C. The block will accommodate one 96-well PCR plate, 96 x 0.2 ml PCR tubes, or 77 x 0.5 ml PCR tubes.
The Eppendorf Mastercycler personal is a thermal cycler designed for the amplification of nucleic acids. This thermal cycler possesses a programmable heated lid and sample block which can accommodate 25 x 0.2 ml PCR tubes, 16 x 0.5 ml PCR tubes or one 25-well PCR plate. Peltier elements control rapid heating speeds up to 3°C per second and cooling speeds up to 2°C per second in a temperature range of 4°C to 99°C.
The GeneAmp® PCR System 9700 is designed for the amplification of nucleic acids. The sample block can accommodate either 96 x 0.2 ml PCR tubes or one 96-well PCR plate. The gold-plated silver block delivers rapid heating and cooling speeds up to 5°C per second, decreasing cycle times for volumes up to 100 µl.
The QIAGEN TissueLyser provides efficient disruption of tissues and cells through high speed shaking and grinding effects of stainless steel beads on biological samples. Up to 48 samples can be disrupted in 2 to 4 minutes, which can then be used to extract RNA, DNA and protein for downstream molecular applications.
The Labconco CentriVap refrigerated concentrator serves to rapidly concentrate small volumes of heat sensitive biological samples, such as RNA, DNA and protein. The CentriVap uses centrifugal force (max rotor speed of 1,725 rpm) coupled with vacuum and a temperature controlled environment ranging from -4°C to 100°C to accelerate the evaporation of up to 132 sample volumes of 1 ml or less.
The Beckman Coulter Optima L-90K is a floor model preparative ultracentrifuge that is able to generate 694,000 x g at speeds up to 90,000 rpm. Available rotors include the SW 41 Ti rotor, with a maximum speed of 41,000 rpm, the SW 55 Ti rotor, with a maximum speed of 55,000 rpm and the Type 50.2 Ti rotor, with a maximum speed of 50,000 rpm, and the Type 90 Ti rotor, with a maximum speed of 90,000 rpm.
The Eppendorf 5810 R is a refrigerated bench top centrifuge. Available rotors include the FA-45-30-11 rotor, which will accommodate 1.5 ml or 2.0 ml microcentrifuge tubes with a maximum speed of 14,000 rpm. The A-4-62 rotor is also available, which will accommodate 15 ml and 50 ml tubes, as well as 96 well plates with a maximum speed of 4,000 rpm.
The Eppendorf 5424 is a non-refrigerated bench top centrifuge that is able to generate speeds up to 15,000 rpm. Two standard aerosol tight rotors are available that will accommodate up to 24 x 1.5 ml or 2.0 ml microcentrifuge tubes, or 8 x 0.2 ml PCR strip tubes.
The UVP CL-1000 Ultraviolet Crosslinker produces shortwave ultraviolet radiation at 254 nm which can be utilized to covalently bind nucleic acids to nitrocellulose, nylon, and nylon-reinforced membranes for Northern, Southern, slot or dot blotting applications. This system can also be used to nick Ethidium Bromide stained DNA in agarose gels for CHEF assays, control the extent of restriction enzyme digestions, detect recA mutations, and sterilize small surfaces by killing bacteria, viruses, bacteriophages and small organisms.
The Bio-Rad DCode Universal Mutation Detection System can be used to screen for DNA polymorphisms. This system provides a controlled temperature environment ranging from 5-70°C and is capable of running up to 64 samples in 2 hours. This system can detect single-base changes through single-stranded conformation polymorphism, denaturing gradient gel electrophoresis, temporal temperature gradient electrophoresis, and constant denaturing gel electrophoresis techniques.
The high capacity VWR hybridization oven functions to meet hybridization needs for Northern, Southern and Western blotting. This oven holds 100, 150, 225 and 300 mm length bottles and can also be adapted as a platform rocker to hold plastic containers or heat sealed bags. This oven is equipped with an adjustable speed carousel and produces three-dimensional motion to provide excellent membrane coverage.
The VWR incubator model 1525 is a general purpose incubator. This incubator is set at 37°C and is without gas or humidity. The temperature range of this unit is 5°C to 70°C.
The UNLV Genomics Core Facility offers all the hardware necessary to run SDS PAGE and Western blots, including single and multi-gel casting systems, rigs, power packs, a gel dryer and a Bio-Rad Bio-Dot SF apparatus.
The UNLV Genomics Core Facility offers all the hardware necessary to run 2D gel electrophoresis.
The UNLV Genomics Core Facility offers all of the hardware needed to run DNA agarose gels, including gel rigs, combs, power packs and a UV illuminator.
The BioRad Gel Dryer Model 583 is designed to dry PAGE, gradient and sequencing gels. The gel dryer contains three preset microprocessor-controlled heating cycles combined with vacuum which provide optimal drying conditions for different gel types. Manual programming allows the user to select any drying temperature between 50°C and 90°C. Gels can be dried on either filter paper or cellophane membranes.
The Techne TC-5000 is a touch screen gradient thermal cycler designed for the amplification of nucleic acids. This thermal cycler has a programmable heated lid and gradient function, which allows the entire block temperature to be varied within a 30°C range. The TC-5000 has a typical heating rate of 3°C per second and cooling rate of 1.3°C per second in a temperature range of 4°C to 99°C. The block will accommodate one 96-well PCR plate or 96 x 0.2 ml PCR tubes.
The Lab-Line Environette Environmental Chamber can reach temperatures up to 60 degrees Celsius. The interior dimensions are approximately 35 inches wide by 24 inches deep by 66 inches high. A power outlet is available inside of the chamber.
Supply Centers
The Invitrogen Supply Center stocks various research-oriented products for purchase on-site, allowing for immediate procurement of these products. Shipping costs are waived for products purchased from the Supply Center. Orders are placed through the PI’s online account. Stocked items are available for immediate pick up and a replenishment is automatically shipped to maintain the Supply Center stock. Users can request items to be added to the onsite stock. Users have the option to ship non-stocked products to the Supply Center without paying shipping fees.